Overview
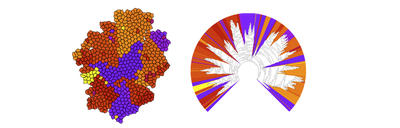
When modeling stem cells, development or tumor progression, we are often interested in the divisional history of cells and their progeny. This can, for example, provide information on the state of differentiation of a cell lineage or the clonal heterogeneity in a tissue.
Morpheus provides a number of ways to track cells and record their division history. Combined with the power of tools to visualize and analyse phylogenetic tree such as ETE3 (Heurta-Capas et al., 2016), this allows for in-depth analysis of cellular genealogies.
In this tutorial, we will show:
- How to export the divisional history from Morpheus using the
CellDivisionplugin. - How to visualize genealogical trees using the python package ETE3, and
- How to execute shell scripts during a Morpheus simulation using the
Externalplugin.